My name is tokibi, and I am mainly in charge of machine learning infrastructure maintenance at COGNANO, a company that integrates biotechnology and IT. In this article, I would like to talk about what I am working on in improving the workflow of machine learning, as well as introduce what kind of work an IT engineer like me, who has built his career around software development, is doing at COGNANO.
Machine Learning Efforts at COGNANO
COGNANO is working on an attempt to build a deep learning model to predict antigen-antibody interactions by generating teacher data based on information about VHH antibodies produced by alpacas. the bioteam has been collecting data for many years since 2010, when the potential of VHH antibodies was first noticed The team has been collecting data for many years. The information reflecting the immune response of the alpaca organism has amassed enough data to be applied to machine learning, with some 300 million records currently in the data set. For a more detailed explanation of machine learning, my colleague Tsurubei, who is in charge of research and development, has published an article on the subject.
Machine Learning Initiatives with Alpaca Antibody and Future
MLOps in Practice.
The IT team should also support machine learning engineers and data scientists through the development of the environment so that they can smoothly perform tasks that lead to improved model performance.
On the introduction of MLOps
The concept of MLOps has emerged in recent years as a practical method for improving the overall efficiency of the workflow in machine learning, and COGNANO is in the process of trying to improve each process while following these principles.
MLOps is a comprehensive concept that covers everything from data collection and preprocessing, which forms the basis of teacher data, to deployment of models and monitoring after the start of operation, etc. This time, we would like to focus on one part of the process, data preprocessing, and discuss our actual efforts.
Assumptions for data preprocessing for VHH antibodies
Although we have just said in a few words that we will create teacher data based on the information of VHH antibodies, several steps must be taken in order to make the data available for machine learning. There are two major processes involved in this: the first half is the processing process for the output of the next-generation sequencer, and the second half is the process of assigning correct labels to pairs of amino acid sequences of the antigen and the VHH antibody.
Antibody genes collected from alpacas are subjected to next-generation sequencing through experiments performed by the bioteam, and the final output is a file described in FASTQ format. This contains the DNA sequences read and the quality score (certainty) of each sequence letter.
Since the output FASTQ file may contain unnecessary sequences or sequences of low quality, it is necessary to perform some initial processing such as adapter trimming and quality trimming, or to convert the nucleotide sequence to an amino acid sequence for subsequent processing. These are combined with several command line tools used in bioinformatics, such as cutadapt and Trimmomatic. They are realized by combining several command line tools used in the field of bioinformatics, such as cutadapt and Trimmomatic, and are basically pipelined processes in which inputs and outputs are connected by commands.
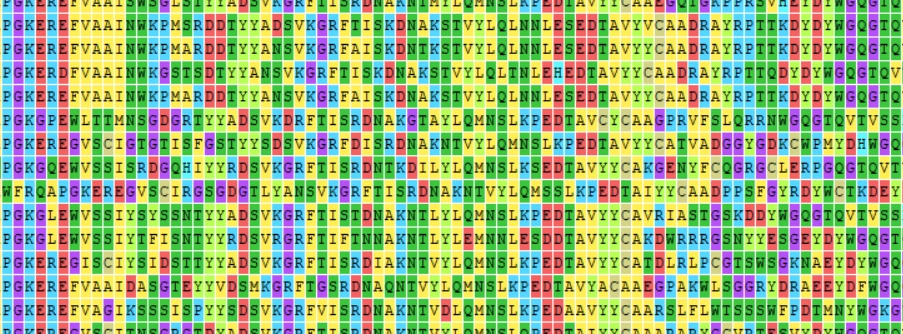
COGNANO uses a proprietary algorithm to compare multiple amino acid sequences generated by biopanning to verify the presence or absence of antigen-antibody binding and labeling. COGNANO performs labeling by comparing multiple amino acid sequence groups generated by biopanning to verify the presence or absence of antigen-antibody binding. The details of the labeling process are beyond the scope of this article, but will be explained in a future paper to be submitted.
Current Issues and Future Efforts
The data preprocessing process we have discussed so far has been automated to some extent in some parts, but some parts of the labeling are done manually using spreadsheet tools. In addition, each time new antibody information is added through experiments, data preprocessing is performed, teacher data including past information is created, and the performance of the model created through training is evaluated. The following two issues have emerged from this process and will be addressed in the future. 1.
- integration of domain knowledge and IT technology
- acquiring domain knowledge of biotechnology by engineers
1. integration of domain knowledge and information technology
In addition to the overall workflow issues mentioned earlier, there is also room for improvement in the parameters and methods used within individual processes. Currently, the selection is based on the domain knowledge of the bio-team, but it is quite possible that better options exist, and an exhaustive comparative study will be necessary in the future. To improve these issues, we are currently starting with the following basic environment based on the principles of DevOps, the predecessor of MLOps
- Using cloud storage for data storage so that necessary processes can be called up by triggering events such as uploads
- Containerizing the execution environment to improve reproducibility and portability
- Build a pipeline for workflow automation
In the future, it will also be necessary to address processes specific to machine learning, such as the development of a model development environment and the evaluation of the impact of parameters and methods on model performance. To this end, we are also looking at introducing open source projects such as Kubeflow and products that comprehensively support the execution of MLOps, which are provided by mega-cloud companies.
In introducing such products, we believe that in some cases it will be easier to iterate on modifications by moving processing to Python or other code. Currently, we are taking the option of containerizing the execution environment for the parts that depend on multiple command line tools, but we plan to move forward with the migration of execution methods as needed.
2. engineers to acquire domain knowledge of biotechnology
In addition, I believe that it will be necessary to acquire knowledge of the biotech domain. I myself am an engineer who has focused on software development in the past, so I do not have expert knowledge in the fields of biotechnology or bioinformatics. Therefore, as we work to improve each process, there are many areas where we do not understand the intent of the process. Fortunately, we have an environment where we can easily ask questions to experts both synchronously and asynchronously via web conferencing and Slack, so I am not troubled by this. However, in order to improve the quality of details and build a better infrastructure environment, I will need to acquire a certain level of domain knowledge myself.
COGNANO's values include "cross-border and fusion," and we place great importance on respecting each other's expertise, while at the same time being as close as possible. This is because we believe that both sides are trying to transcend the bio and IT domains, and the results that emerge from this will create great value. The construction of the deep learning model mentioned in this article is one of the results of these efforts. Through my ongoing projects, I would like to work every day to make a better contribution to society.
Conclusion
In this article, I introduced the pre-processing process of machine learning at COGNANO and our efforts to apply it to the MLOps framework. Although I interjected some hard talk of social contribution at the end, I myself enjoy learning more and more about unknown areas and working on improvements every day.
I plan to publish details of the data sets we are currently using and the pre-processing in conjunction with the submission of papers and other activities. We hope to be able to provide the data with as high a level of reproducibility as possible, so if you are interested in our work, please look forward to it.
Thank you for reading to the end, and if you have anything to say about COGNANO, please contact CEO Imura, CTO Matsumotory and tokibi.